Enumerating metabolic pathways for the production of heterologous target chemicals in chassis organisms, BMC Systems Biology
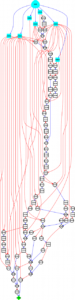
When considering the best metabolic pathways to engineer into a chassis organism in order to synthesize heterologous target compounds, the problem of enumerating all possible heterologous pathways linking a target compounds to a source set of compounds needs to be addressed. Following our previous work on a retrosynthetic biology approach that searches the metabolic space, we explore in this paper the performance of two methods based on steady-state analysis and hypergraph network topology. This work, thus, provides a powerful tool for pathway enumeration with direct application to biosynthetic pathway design.
For instance, there exists at least 224 possible metabolic pathways for producing typhasterol, a heterologous compound in E. coli. The number however, can become significantly larger if additional putative promiscuous enzymatic reactions and supplements are considered, making necessary the use of a tool such as the one that we have developed to efficiently enumerate all possible pathways.
Carbonell, P, Fichera, D, Pandit, SB, Faulon, JL. Enumerating metabolic pathways for the production of heterologous target chemicals in chassis organisms. BMC Systems Biology, in press, 2012.



