Protein function prediction
We have developed a new technique to predict protein (enzymatique) function, protein-protein and protein-chemical interactions using the signature descriptor described in the QSAR pages. The signature of a protein sequence (or DNA sequence) is a vector of occurence numbers of the sequence k-mers (or k-words). To predict protein binding pairs a new type of kernel was developed where the signagture is a product of the signatures of both the bait (target) and the prey (ligand). The kernel, which is used with SVM compares well in term of accuracy/precision/sensitivity with other existing techniques. Our method is not specific to a particular type of experimental dataset, does not require knowledge of protein domains or physico-chemical parameters, it just uses as input a list of sequences and binding pairs. More information can be found in the following papers:
- Faulon J.L., Misra M., Martin S., Sale, K., Sapra R. Genome Scale Enzyme-metabolites and Drug-Target interaction predictions using the signature molecular descriptor, Bioinformatics in press 2007 [PMID: 18037612 ] (link to journal)
- Martin S., Brown W. M., Faulon J.L. Using Product Kernels to Predict Protein Interactions. In Advances in Biochemical Engineering/Biotechnology, Eds. H. Seitz and M. Werther, Springer-Verlag. in press, 2007 [PMID: 17922100]
- Brown W. M., Martin S., Chabarek J.P., Strauss C, Faulon J.L. Prediction of β-Strand Packing Interactions using the Signature Product, Journal of Molecular Modeling, in press. [PMID: 16365772] (link to journal)
- Martin S., Roe D., Faulon J.L. Predicting Protein-Protein Interactions using Signature Products, Bioinformatics, 21, 218-26, 2005. [PMID: 15319262]. (.pdf manuscript)
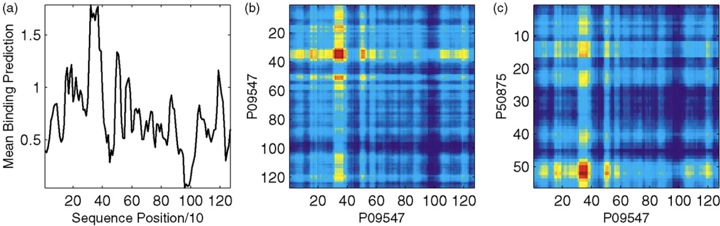
Plot (a) shows the mean binding activity (predicted by signature kernel) of a window (50 residue long) along with x-axis for protein P09547, a Yeast protein involved in transcriptional regulation. Plot (b) shows an intensity plot of the binding activities of all pairs of windows in P09547; and (c) shows an intensity plot of the binding activities of all pairs of windows in P09547 and P50875 (a transcription factor in Yeast). In (b and c), red denotes activity and blue denotes inactivity. An hypothetical domain is found for P09547 between residue 30×10 =300 and residue 400. The domain matches a Gln-rich Swiss-prot domain (residue 337 to 385), as well as the InterPro entry IPR001660 SAM. The domain found for P50875 following residue 50×10=500 matches a Poly-Asn Swiss-prot domain (residue 552 to 559).



