BioRetroSynth: Metabolic Engineering using Retrosynthesis
Retrosynthesis graphs for heterologous antibiotics production in E. coli
Synthetic biology  addresses problems beyond classical metabolic engineering by importing pathways from other organisms into microbial chassis. In this project we are focusing into production of therapeutics belonging to the family of antimicrobial agents with the goal of developing an in situ drug delivery device in host cells. The process consists of implementing a heterologous circuit in E. coli by using retrosynthesis, a concept originally developed for synthetic chemistry.
addresses problems beyond classical metabolic engineering by importing pathways from other organisms into microbial chassis. In this project we are focusing into production of therapeutics belonging to the family of antimicrobial agents with the goal of developing an in situ drug delivery device in host cells. The process consists of implementing a heterologous circuit in E. coli by using retrosynthesis, a concept originally developed for synthetic chemistry.
Retrosynthesis consists of applying iteratively reversed biotransformations (i.e. reversed enzymes-catalyzed reactions) starting from a target product, in order to reach precursors that are metabolites endogenous to the host organism. Potential pathways are constructed by using a reaction chemical similarity search based on molecular signatures in order to predict reactions producing the target compound or intermediates. Candidate pathways are then ranked to select which pathways are best to engineer. The ranking function is based on several criteria such as inhibitory effects, cytotoxicity of heterologous metabolites, and host compatibility (codon usage, homology). Furthermore, we use several predictive tools (the MolSig package) developed by our group in order to estimate Quantitative Structure-Activity Relationships (QSAR) for enzyme activity and reaction efficiency at each step of the identified pathways. Based on these in silico predictions, exogenous pathways are improved by selecting enzyme variants with enhanced performance.
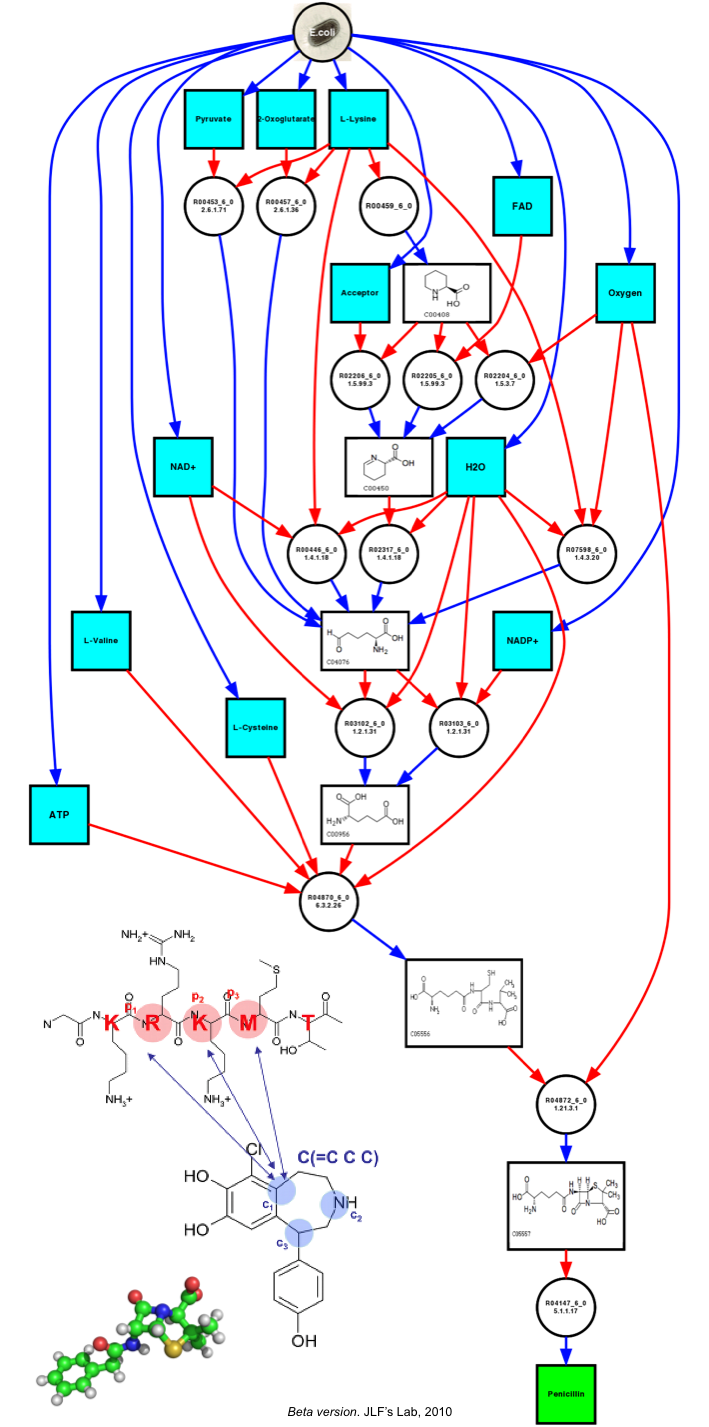
For instance, we provide here an example of the retrosynthesis map generated by our algorithm for penicillin, starting from a given set of reaction constraints based on the KEGG database of metabolic pathways. The algorithm found 15 pathways involving the insertion of 5 up to 7 heterologous genes.



